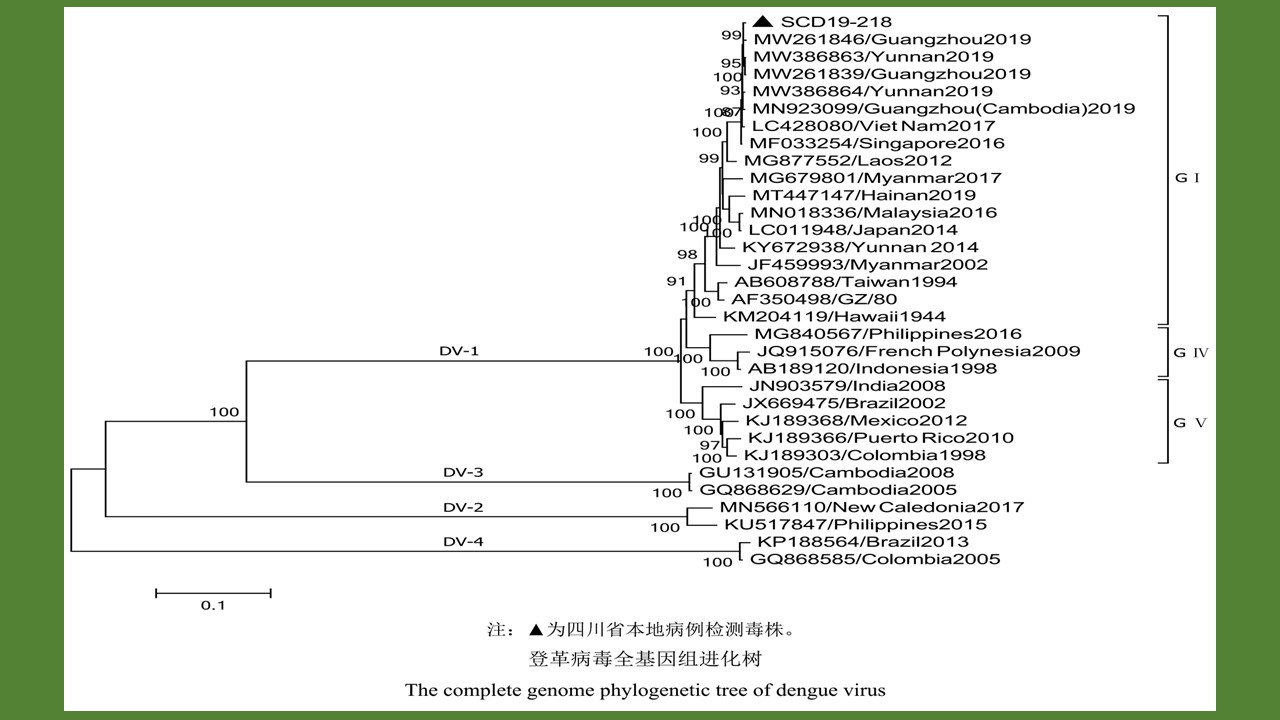
目的 对2019年四川省发生的首起本地传播登革热疫情中检出的登革病毒进行全基因组测序,分析病毒分子生物学特征并探讨其可能的来源。方法 对实时荧光定量PCR(RT-qPCR)检测为阳性的本地病例标本提取核酸并扩增全基因组序列和测序,测序后进行同源性、系统进化分析和变异分析。结果 通过基因扩增和序列测定获得1株病毒全基因组序列和2株包括E基因的全基因组部分序列,全基因组序列全长10 706 bp,编码3 393个氨基酸;同源性和系统进化分析显示,本次疫情流行株与2019年我国广州市分离株、2019年广州市分离的柬埔寨输入株、2019年云南省分离株、2017年越南分离株以及2016年新加坡分离株的同源性较高,系统进化属于同一分支;与登革1型原型株相比本次流行株散在分布有589个核苷酸变异和64个氨基酸突变,但影响毒力的主要氨基酸位点没有改变。结论 2019年四川省首起登革热本地暴发疫情可能由柬埔寨输入病例引起,其病原体为登革1型病毒基因Ⅰ型;本次四川省流行株的毒力相关基因没有明显改变,但其在某些相关区域发生点突变的生物学意义有待进一步研究。
Objective To sequence the complete genome of Dengue virus from the first local dengue outbreak in Sichuan province, China, 2019, and to analyze its molecular biological characteristics and possible origin. Methods Nucleic acids were extracted and whole genome sequences were amplified and sequenced from local case specimens that tested positive by real-time PCR, followed by homology analysis, phylogenetic analysis, and variation analysis. Results The complete genome sequence of one virus strain and the partial genome sequences (containing the E gene) of two virus strains were obtained. The complete genome was 10 706 bp long, encoding 3 393 amino acids. Homology and phylogenetic analyses showed that the epidemic strain in Sichuan province had high homology to the Guangzhou (China) strain isolated in 2019, the Guangzhou strain imported from Cambodia in 2019, the Yunnan province (China) strains isolated in 2019, the Vietnam strain isolated in 2017, and the Singapore strain isolated in 2016, belonging to the same branch in the phylogenetic tree. Compared with the prototype strain of Dengue virus type 1, the Sichuan strain had 589 nucleotide variations and 64 amino acid mutations that were scattered, but no alteration occurred at the key amino acid sites affecting virulence. Conclusion The first local dengue fever outbreak in Sichuan province, 2019 may be caused by imported cases from Cambodia, and the pathogen was the Dengue virus type 1, genotypeⅠ. The strain of Dengue virus isolated from Sichuan province has no marked alteration in virulence-related genes, but the biological significance of the point mutations in some sites remains to be further studied.
[1] 张拥军, 吴生根, 王金章, 等. 福建省2015年本地登革热病例的病原学特征[J]. 中国人兽共患病学报, 2019, 35(1):28-33. DOI:10.3969/j.issn.1002-2694.2018.00.210. Zhang YJ, Wu SG, Wang JZ, et al. Etiological characterization of indigenous dengue cases in Fujian province, 2015[J]. Chin J Zoonoses, 2019, 35(1):28-33. DOI:10.3969/j.issn.1002-2694. 2018.00.210.(in Chinese)
[2] 刘起勇. 新时代媒介生物传染病形势及防控对策[J]. 中国媒介生物学及控制杂志, 2019, 30(1):1-6, 11. DOI:10.11853/j.issn.1003.8280.2019.01.001. Liu QY. Epidemic profile of vector-borne diseases and vector control strategies in the new era[J]. Chin J Vector Biol Control, 2019, 30(1):1-6, 11. DOI:10.11853/j.issn.1003.8280.2019.01. 001.(in Chinese)
[3] 刘永华, 尹小雄, 张海林, 等. 云南省德宏州2013-2019年登革热流行特征及媒介伊蚊监测分析[J]. 中国媒介生物学及控制杂志, 2021, 32(2):173-180. DOI:10.11853/j.issn.1003. 8280.2021.02.011. Liu YH, Yin XX, Zhang HL, et al. Epidemiological characteristics of dengue fever and monitoring of Aedes vector mosquitoes in Dehong Dai and Jingpo Autonomous Prefecture of Yunnan province, China, 2013-2019[J]. Chin J Vector Biol Control, 2021, 32(2):173-180. DOI:10.11853/j.issn.1003. 8280.2021.02.011.(in Chinese)
[4] 刘媛媛, 刘远, 罗雷, 等. 2011-2019年广州市登革热流行病学特征分析[J]. 现代预防医学, 2021, 48(11):1925-1929. Liu YY, Liu Y, Luo L, et al. Epidemiological analysis on dengue fever cases in Guangzhou, 2011-2019[J]. Mod Prev Med, 2021, 48(11):1925-1929. (in Chinese)
[5] Wu WL, Bai ZJ, Zhou HQ, et al. Molecular epidemiology of dengue viruses in southern China from 1978 to 2006[J]. Virol J, 2011, 8(1):322. DOI:10.1186/1743-422X-8-322.
[6] 李杨, 张文宏. 全球登革热疫情态势、疫情警报[J]. 中华传染病杂志, 2019, 37(10):619-621. DOI:10.3760/cma.j.issn.1000-6680.2019.10.008. Li Y, Zhang WH. Global dengue epidemic situation and alert[J]. Chin J Infect Dis, 2019, 37(10):619-621. DOI:10.3760/cma.j.issn.1000-6680.2019.10.008.(in Chinese)
[7] 马珍元, 黄呈辉, 张新枝, 等. 2014年深圳市流行的1型登革病毒分子溯源分析[J]. 中国现代医学杂志, 2016, 26(3):39-44. DOI:10.3969/j.issn.1005-8982.2016.03.008. Ma ZY, Huang CH, Zhang XZ, et al. Molecular phylogenetic analysis of type 1 strains of dengue virus isolated in Shenzhen city during the 2014 dengue outbreak[J]. China J Mod Med, 2016, 26(3):39-44. DOI:10.3969/j.issn.1005-8982.2016.03. 008.(in Chinese)
[8] Tamura K, Stecher G, Peterson D, et al. MEGA 6.0:Molecular evolutionary genetics analysis version 6.0[J]. Mol Biol Evol, 2013, 30(12):2725-2729. DOI:10.1093/molbev/mst197.
[9] Guirakhoo F, Zhang Z, Myers G, et al. A single amino acid substitution in the envelope protein of chimeric yellow fever-dengue 1 vaccine virus reduces neurovirulence for suckling mice and viremia/viscerotropism for monkeys[J]. J Virol, 2004, 78(18):9998-10008. DOI:10.1128/JVI.78.18.9998-10008.2004.
[10] 赵卫, 杨佩英. 登革病毒毒力研究进展[J]. 中华实验和临床病毒学杂志, 2001, 15(2):197-199. DOI:10.3760/cma.j.issn. 1003-9279.2001.02.037. Zhao W, Yang PY. Research progress on virulence of dengue virus[J]. Chin J Exp Clin Virol, 2001, 15(2):197-199. DOI:10.3760/cma.j.issn.1003-9279.2001.02.037.(in Chinese)
[11] Holmes EC, Worobey M, Rambaut A. Phylogenetic evidence for recombination in dengue virus[J]. Mol Biol Evol, 1999, 16(3):405-409. DOI:10.1093/oxfordjournals.molbev.a026121.
[12] Dos Santos CND, Frenkiel MP, Courageot MP, et al. Determinants in the envelope E protein and viral RNA helicase NS3 that influence the induction of apoptosis in response to infection with dengue type 1 virus[J]. Virology, 2000, 274(2):292-308. DOI:10.1006/viro.2000.0457.
[13] 曹一鸥, 吕强, 张佳珂, 等. 2019年四川省首例登革热本地病例调查与疫区处置情况分析[J]. 预防医学情报杂志, 2020, 36(11):1499-1503. Cao YO, Lyu Q, Zhang JK, et al. Investigation and disposal of the first local case of dengue fever in Sichuan province[J]. J Prev Med Inf, 2020, 36(11):1499-1503. (in Chinese)
[14] 李晓萸, 耿丽卿, 苑锡同, 等. 我国登革1型病毒广州株基因组全序列测定与分析[J]. 军事医学科学院院刊, 2001, 25(3):161-165. DOI:10.3969/j.issn.1674-9960.2001.03.001. Li XY, Geng LQ, Yuan XT, et al. Determination and analysis of the complete genome of dengue 1 virus GZ/80 strain[J]. Bull Acad Mil Med Sci, 2001, 25(3):161-165. DOI:10.3969/j.issn.1674-9960.2001.03.001.(in Chinese)
[15] Cahour A, Pletnev A, Vazeille-Falcoz M, et al. Growth-restricted dengue virus mutants containing deletions in the 5'noncoding region of the RNA genome[J]. Virology, 1995, 207(1):68-76. DOI:10.1006/viro.1995.1052.
[16] Men R, Bray M, Clark D, et al. Dengue type 4 virus mutants containing deletions in the 3'noncoding region of the RNA genome:Analysis of growth restriction in cell culture and altered viremia pattern and immunogenicity in rhesus monkeys[J]. J Virol, 1996, 70(6):3930-3937. DOI:10.1128/JVI.70.6.3930-3937.1996.



